Assessing Variants in a Known Gene: Clinical Variant Classification and Use of the ClinGen Variant Curation Interface - ClinGen | Clinical Genome Resource

The Clinical Variant Analysis Tool: Analyzing the evidence supporting reported genomic variation in clinical practice - ScienceDirect

Clinical DNA Variant Interpretation: Theory and Practice (Translational and Applied Genomics): 9780128205198: Medicine & Health Science Books @ Amazon.com

Mutliplexed functional data informs clinical variant interpretation — Center for the Multiplex Assessment of Phenotype

The Clinical Variant Analysis Tool: Analyzing the evidence supporting reported genomic variation in clinical practice - ScienceDirect

Risk of severe clinical outcomes among persons with SARS-CoV-2 infection with differing levels of vaccination during widespread Omicron (B.1.1.529) and Delta (B.1.617.2) variant circulation in Northern California: A retrospective cohort study -

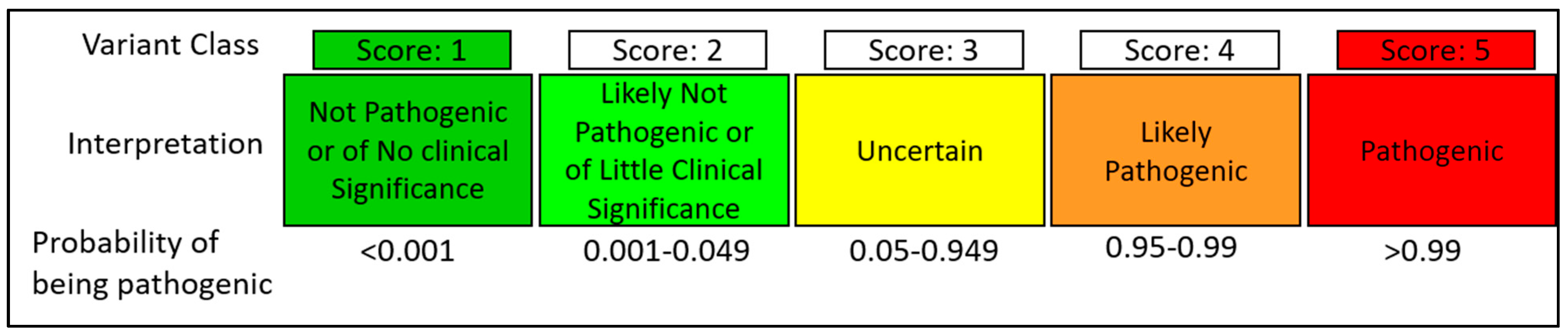
Nothing's for sure, that's for sure: Evaluating variants of uncertain significance | Beyond the Ion Channel
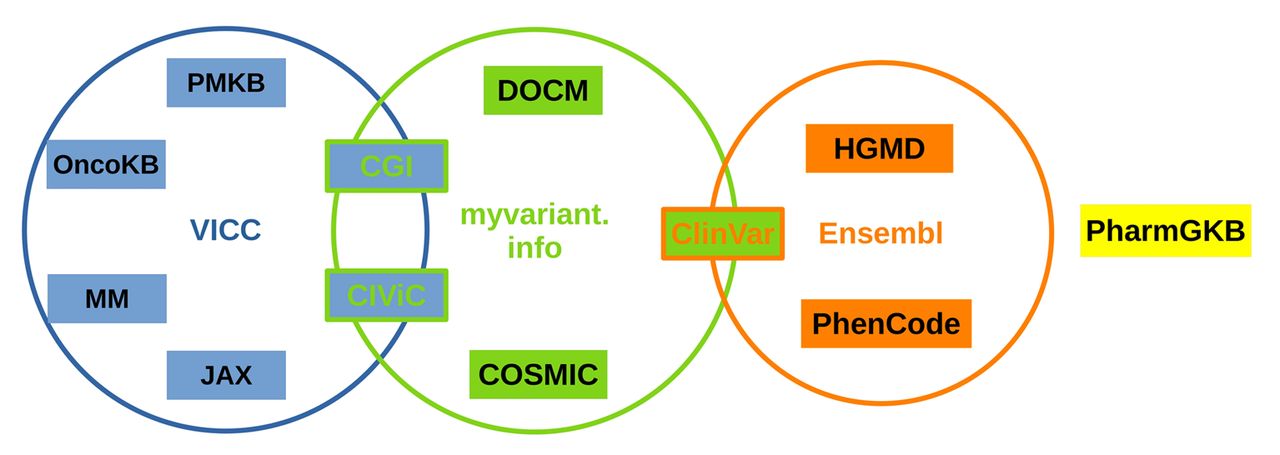
Comparison of Open-access Databases for Clinical Variant Interpretation in Cancer: A Case Study of MDS/AML | Cancer Genomics & Proteomics
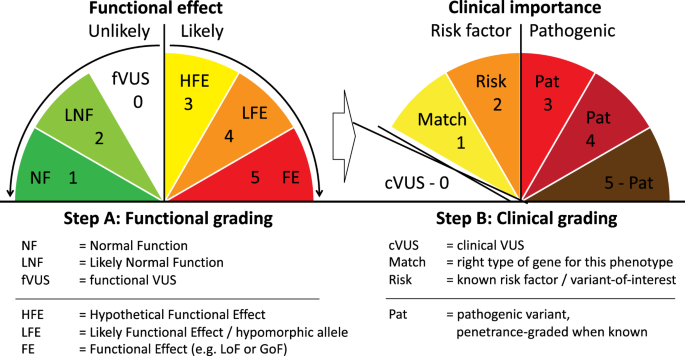
Stepwise ABC system for classification of any type of genetic variant | European Journal of Human Genetics

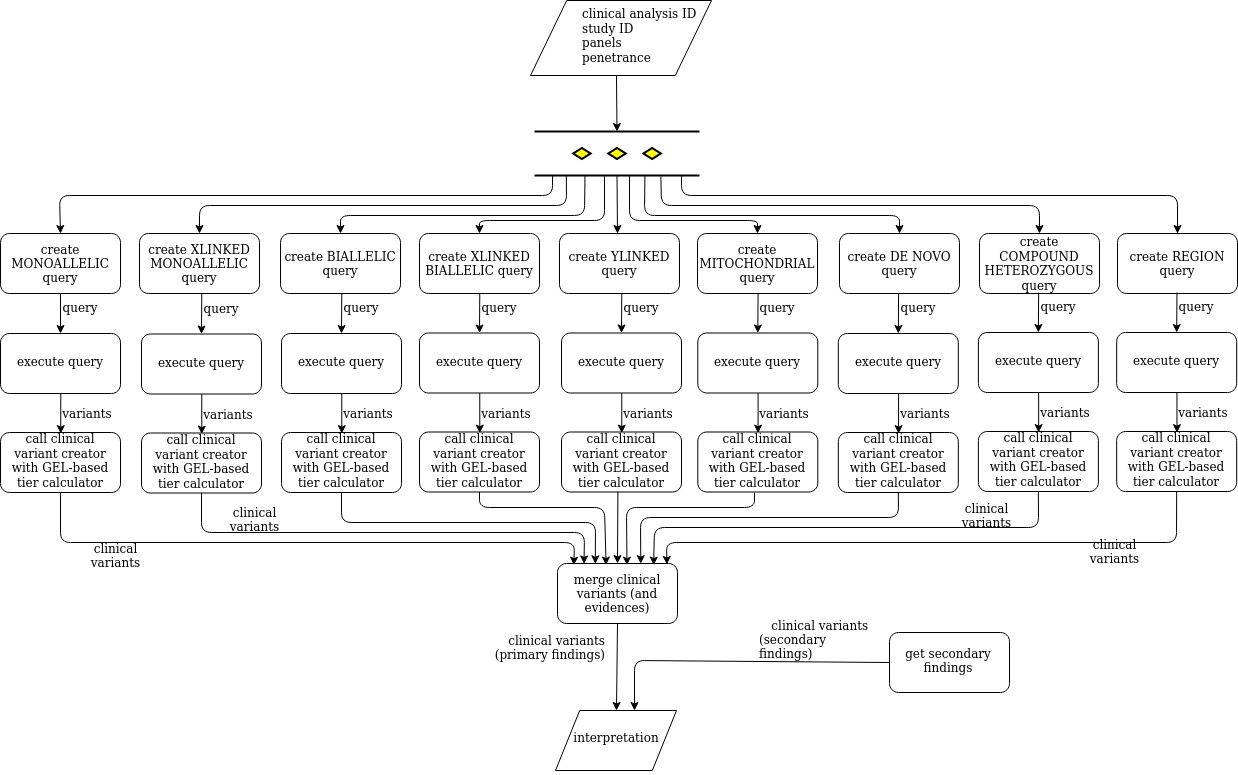
Matching whole genomes to rare genetic disorders: Identification of potential causative variants using phenotype‐weighted knowledge in the CAGI SickKids5 clinical genomes challenge - Pal - 2020 - Human Mutation - Wiley Online Library

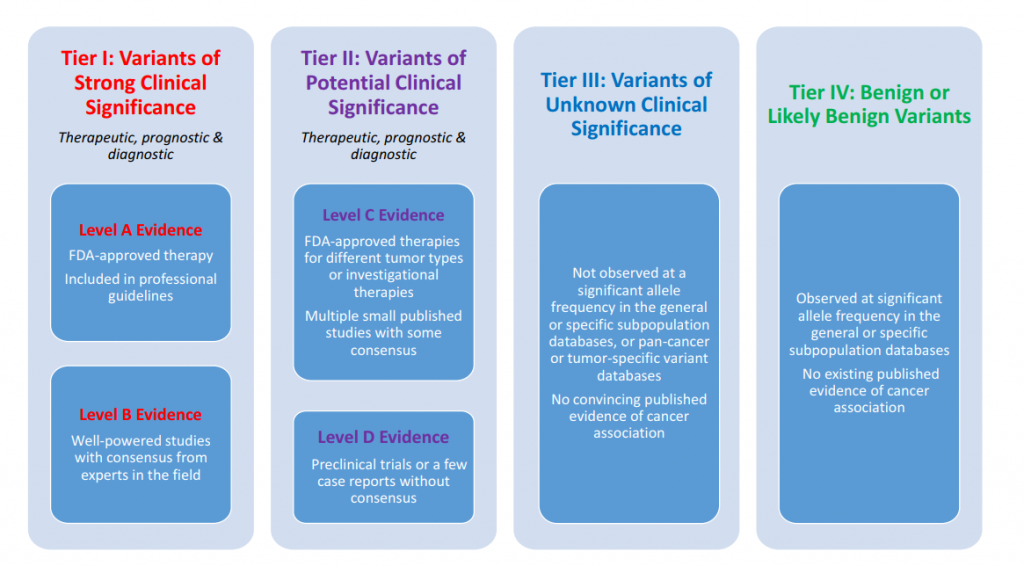
Standards for the classification of pathogenicity of somatic variants in cancer (oncogenicity): Joint recommendations of Clinical Genome Resource (ClinGen), Cancer Genomics Consortium (CGC), and Variant Interpretation for Cancer Consortium (VICC ...

Clinical Variant Classification: A Comparison of Public Databases and a Commercial Testing Laboratory. | Semantic Scholar
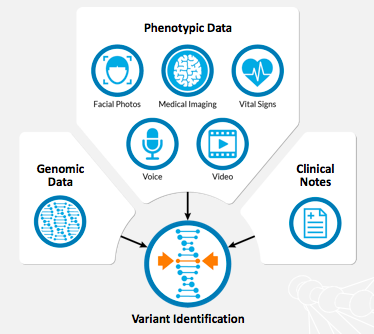
Dr. Christine Stanley: Face2Gene LABS at WuXi NextCODE: Phenotyping for Improved Variant Prioritization - FDNA™

CIViC is a community knowledgebase for expert crowdsourcing the clinical interpretation of variants in cancer | Nature Genetics
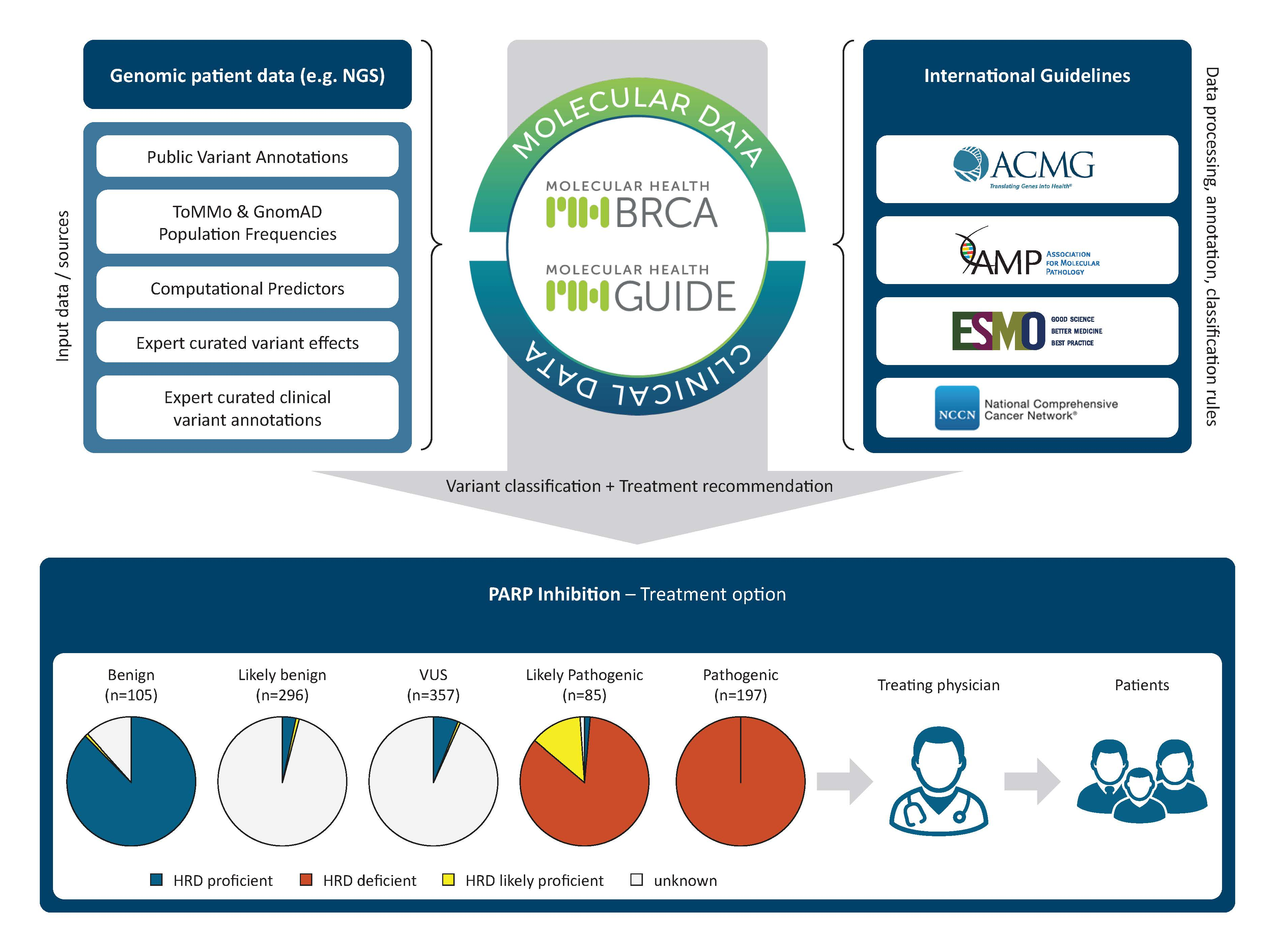
IJMS | Free Full-Text | Consolidated BRCA1/2 Variant Interpretation by MH BRCA Correlates with Predicted PARP Inhibitor Efficacy Association by MH Guide

The ClinGen Sequence Variant Interpretation Working Group: Refining Criteria for Interpreting the Pathogenicity of Genetic Variants -

Genomic Variant Analysis & Clinical Interpretation | Council of Scientific & Industrial Research | CSIR | GoI